Text
Data Management and Visualization_Nguyen Thi Minh Ha_Week 4- Assignment
# -*- coding: utf-8 -*-
"""
Created on Thu Oct 7 00:37:40 2021
@author: hantm
"""
import pandas as pd
import seaborn as sb
import matplotlib.pyplot as plt
# load gapminder dataset
data = pd.read_csv('gapminder.csv',low_memory=False)
# lower-case all DataFrame column names
data.columns = map(str.lower, data.columns)
# bug fix for display formats to avoid run time errors
pd.set_option('display.float_format', lambda x:'%f'%x)
# setting variables to be numeric
data['suicideper100th'] = data['suicideper100th'].apply(pd.to_numeric, errors='coerce')
data['breastcancerper100th'] = data['breastcancerper100th'].apply(pd.to_numeric, errors='coerce')
data['hivrate'] = data['hivrate'].apply(pd.to_numeric, errors='coerce')
data['employrate'] = data['employrate'].apply(pd.to_numeric, errors='coerce')
# display summary statistics about the data
# print("Statistics for a Suicide Rate")
# print(data['suicideper100th'].describe())
# subset data for a high suicide rate based on summary statistics
sub = data[(data['suicideper100th']>12)]
#make a copy of my new subsetted data
sub_copy = sub.copy()
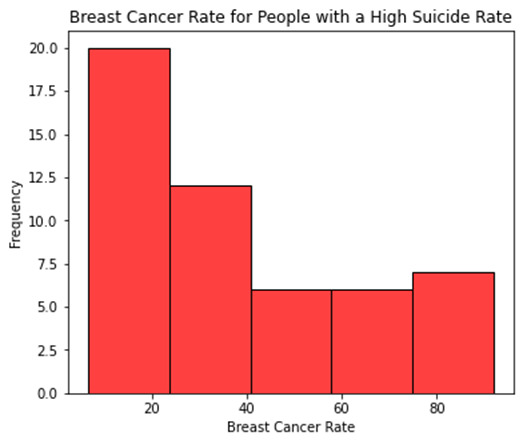
# Univariate graph for breast cancer rate for people with a high suicide rate
plt.figure(1)
sb.histplot(sub_copy["breastcancerper100th"], color="red", kde=False, bins=5)
#sb.distplot(sub_copy["breastcancerper100th"].dropna(),kde=False)
#sb.histplot(sub_copy["breastcancerper100th"], color="red", label="100% Equities", kde=True, stat="density", linewidth=0)
#sb.histplot(sub_copy["hivrate"],kde=False)
plt.xlabel('Breast Cancer Rate')
plt.ylabel('Frequency')
plt.title('Breast Cancer Rate for People with a High Suicide Rate')
plt.show()
# Univariate graph for hiv rate for people with a high suicide rate
plt.figure(2)
#sb.distplot(sub_copy["hivrate"].dropna(),kde=False)
#sb.histplot(sub_copy["hivrate"], color="red", label="100% Equities", kde=True, stat="density", linewidth=0)
sb.histplot(sub_copy["hivrate"], color="red", label="100% Equities", kde=False, stat="density", linewidth=0)
plt.xlabel('HIV Rate')
plt.ylabel('Frequency')
plt.title('HIV Rate for People with a High Suicide Rate')
plt.show()
# Univariate graph for employment rate for people with a high suicide rate
plt.figure(3)
sb.histplot(sub_copy["employrate"], color="red", label="100% Equities", kde=True, stat="density", linewidth=0)
plt.xlabel('Employment Rate')
plt.ylabel('Frequency')
plt.title('Employment Rate for People with a High Suicide Rate')
plt.show()
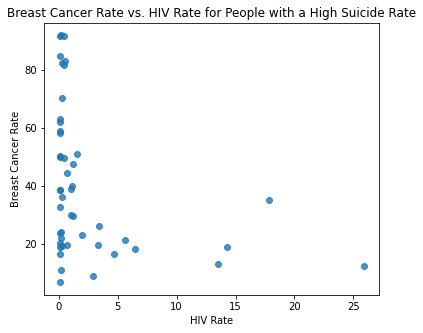
# Bivariate graph for association of breast cancer rate with HIV rate for people with a high suicide rate
plt.figure(4)
sb.regplot(x="hivrate",y="breastcancerper100th",fit_reg=False,data=sub_copy)
plt.xlabel('HIV Rate')
plt.ylabel('Breast Cancer Rate')
plt.title('Breast Cancer Rate vs. HIV Rate for People with a High Suicide Rate')
plt.show()
Please see my output file and code running in Python



0 notes
Text
Data Management and Visualization_Nguyen Thi Minh Ha_Week 3- Assignment
# -*- coding: utf-8 -*-
"""
Created on Thu Oct 7 00:37:40 2021
@author: hantm
"""
import pandas as pd
import numpy as np
import seaborn as sb
import matplotlib.pyplot as plt
# load gapminder dataset
data = pd.read_csv('gapminder.csv',low_memory=False)
# lower-case all DataFrame column names
data.columns = map(str.lower, data.columns)
# bug fix for display formats to avoid run time errors
pd.set_option('display.float_format', lambda x:'%f'%x)
# setting variables to be numeric
data['suicideper100th'] = data['suicideper100th'].apply(pd.to_numeric, errors='coerce')
data['breastcancerper100th'] = data['breastcancerper100th'].apply(pd.to_numeric, errors='coerce')
data['hivrate'] = data['hivrate'].apply(pd.to_numeric, errors='coerce')
data['employrate'] = data['employrate'].apply(pd.to_numeric, errors='coerce')
# display summary statistics about the data
# print("Statistics for a Suicide Rate")
# print(data['suicideper100th'].describe())
# subset data for a high suicide rate based on summary statistics
sub = data[(data['suicideper100th']>12)]
#make a copy of my new subsetted data
sub_copy = sub.copy()
# Univariate graph for breast cancer rate for people with a high suicide rate
plt.figure(1)
sb.distplot(sub_copy["breastcancerper100th"].dropna(),kde=False)
plt.xlabel('Breast Cancer Rate')
plt.ylabel('Frequency')
plt.title('Breast Cancer Rate for People with a High Suicide Rate')
# Univariate graph for hiv rate for people with a high suicide rate
plt.figure(2)
sb.distplot(sub_copy["hivrate"].dropna(),kde=False)
plt.xlabel('HIV Rate')
plt.ylabel('Frequency')
plt.title('HIV Rate for People with a High Suicide Rate')
# Univariate graph for employment rate for people with a high suicide rate
plt.figure(3)
sb.distplot(sub_copy["employrate"].dropna(),kde=False)
plt.xlabel('Employment Rate')
plt.ylabel('Frequency')
plt.title('Employment Rate for People with a High Suicide Rate')
# Bivariate graph for association of breast cancer rate with HIV rate for people with a high suicide rate
plt.figure(4)
sb.regplot(x="hivrate",y="breastcancerper100th",fit_reg=False,data=sub_copy)
plt.xlabel('HIV Rate')
plt.ylabel('Breast Cancer Rate')
plt.title('Breast Cancer Rate vs. HIV Rate for People with a High Suicide Rate')
# --------Output file -----------
#runfile('D:/HaNTM/2021/Course_DM/GapMinder/W3_HaNTM.py', #wdir='D:/HaNTM/2021/Course_DM/GapMinder')
Statistics for a Suicide Rate
count 191.000000
mean 9.640839
std 6.300178
min 0.201449
25% 4.988449
50% 8.262893
75% 12.328551
max 35.752872
Name: suicideper100th, dtype: float64
Number of Breast Cancer Cases with a High Suicide Rate
# of Cases Freq. Percent Cum. Freq. Cum. Percent
(0.0, 23.0] 18 33.96 18 33.96
(23.0, 46.0] 15 28.30 33 62.26
(46.0, 69.0] 10 18.87 43 81.13
(69.0, 92.0] 8 15.09 51 96.23
nan 2 3.77 53 100.00
HIV Rate with a High Suicide Rate
Rate Freq. Percent Cum. Freq. Cum. Percent
0% tile 18 33.96 18 33.96
25% tile 8 15.09 26 49.06
50% tile 11 20.75 37 69.81
75% tile 12 22.64 49 92.45
nan 4 7.55 53 100.00
Employment Rate with a High Suicide Rate
Rate Freq. Percent Cum. Freq. Cum. Percent
1 10 18.87 10 18.87
2 24 45.28 34 64.15
3 5 9.43 39 73.58
4 13 24.53 52 98.11
5 1 1.89 53 100.00
#---------------------------------------------------------------------------------------
#p/s: Nguyen Thi Minh Ha (HaNTM)
1 note
·
View note
Text
Data Management and Visualization_Nguyen Thi Minh Ha_Week 2- Assignment 1
---------------------------------------------------------------------------------------------------------
Summary of Frequency Distributions
------------------------------------------------------
Question 1: What is a number of breast cancer cases associated with a high suicide rate?
The high suicide rate is associated with the low number of breast cancer cases.
------------------------------------------------------
Question 2: How HIV rate is associated with a high suicide rate?
The high suicide rate is associated with the low HIV rate.
------------------------------------------------------
Question 3: How employment rate is associated with a high suicide rate?
The high suicide rate occurs at 55% of employment rate.
-----------------------------------------------------------------------------------------------------------
# -*- coding: utf-8 -*-
"""
Created on Tue Oct 5 12:16:40 2021
@author: hantm
"""
import pandas as pd
# load gapminder dataset
data = pd.read_csv('gapminder.csv',low_memory=False)
# lower-case all DataFrame column names
data.columns = map(str.lower, data.columns)
# bug fix for display formats to avoid run time errors
pd.set_option('display.float_format', lambda x:'%f'%x)
# setting variables to be numeric
data['suicideper100th'] = data['suicideper100th'].apply(pd.to_numeric, errors='coerce')
data['breastcancerper100th'] = data['breastcancerper100th'].apply(pd.to_numeric, errors='coerce')
data['hivrate'] = data['hivrate'].apply(pd.to_numeric, errors='coerce')
data['employrate'] = data['employrate'].apply(pd.to_numeric, errors='coerce')
# display summary statistics about the data
print("Statistics for a Suicide Rate")
print(data['suicideper100th'].describe())
# subset data for a high suicide rate based on summary statistics
sub = data[(data['suicideper100th']>12)]
#make a copy of my new subsetted data
sub_copy = sub.copy()
#print(sub)
# BREAST CANCER RATE
# frequency and percentage distritions for a number of breast cancer cases with a high suicide rate
#print('frequency for a number of breast cancer cases with a high suicide rate')
bc = sub_copy['breastcancerper100th'].value_counts(sort=False,bins=10)
#print(bc)
print('percentage for a number of breast cancer cases with a high suicide rate')
pbc = sub_copy['breastcancerper100th'].value_counts(sort=False,bins=10,normalize=True)*100
#print(pbc)
# cumulative frequency and cumulative percentage for a number of breast cancer cases with a high suicide rate
bc1=[] # Cumulative Frequency
pbc1=[] # Cumulative Percentage
cf=0
cp=0
for freq in bc:
cf=cf+freq
bc1.append(cf)
pf=cf*100/len(sub_copy)
pbc1.append(pf)
#print('cumulative frequency for a number of breast cancer cases with a high suicide rate')
#print(bc1)
#print('cumulative percentage for a number of breast cancer cases with a high suicide rate')
#print(pbc1)
print('Number of Breast Cancer Cases with a High Suicide Rate')
fmt1 = '%s %7s %9s %12s %12s'
fmt2 = '%5.2f %10.d %10.2f %10.d %12.2f'
print(fmt1 % ('# of Cases','Freq.','Percent','Cum. Freq.','Cum. Percent'))
for i, (key, var1, var2, var3, var4) in enumerate(zip(bc.keys(),bc,pbc,bc1,pbc1)):
#print(key.left)
print(fmt2 % (key.left, var1, var2, var3, var4))
fmt3 = '%5s %10s %10s %10s %12s'
print(fmt3 % ('NA', '2', '3.77', '53', '100.00'))
# HIV RATE
# frequency and percentage distritions for HIV rate with a high suicide rate
#print('frequency for HIV rate with a high suicide rate')
hc = sub_copy['hivrate'].value_counts(sort=False,bins=7)
#print(hc)
#print('percentage for HIV rate with a high suicide rate')
phc = sub_copy['hivrate'].value_counts(sort=False,bins=7,normalize=True)*100
#print(phc)
# cumulative frequency and cumulative percentage for HIV rate with a high suicide rate
hc1=[] # Cumulative Frequency
phc1=[] # Cumulative Percentage
cf=0
cp=0
for freq in bc:
cf=cf+freq
hc1.append(cf)
pf=cf*100/len(sub_copy)
phc1.append(pf)
#print('cumulative frequency for HIV rate with a high suicide rate')
#print(hc1)
#print('cumulative percentage for HIV rate with a high suicide rate')
#print(phc1)
print('HIV Rate with a High Suicide Rate')
fmt1 = '%5s %12s %9s %12s %12s'
fmt2 = '%5.2f %10.d %10.2f %10.d %12.2f'
print(fmt1 % ('Rate','Freq.','Percent','Cum. Freq.','Cum. Percent'))
for i, (key, var1, var2, var3, var4) in enumerate(zip(hc.keys(),hc,phc,hc1,phc1)):
print(fmt2 % (key.left, var1, var2, var3, var4))
fmt3 = '%5s %10s %10s %10s %12s'
print(fmt3 % ('NA', '2', '3.77', '53', '100.00'))
# EMPLOYMENT RATE
# frequency and percentage distritions for employment rate with a high suicide rate
#print('frequency for employment rate with a high suicide rate')
ec = sub_copy['employrate'].value_counts(sort=False,bins=10)
#print(ec)
#print('percentage for employment rate with a high suicide rate')
pec = sub_copy['employrate'].value_counts(sort=False,bins=10,normalize=True)*100
#print(pec)
# cumulative frequency and cumulative percentage for employment rate with a high suicide rate
ec1=[] # Cumulative Frequency
pec1=[] # Cumulative Percentage
cf=0
cp=0
for freq in bc:
cf=cf+freq
ec1.append(cf)
pf=cf*100/len(sub_copy)
pec1.append(pf)
#print('cumulative frequency for employment rate with a high suicide rate')
#print(ec1)
#print('cumulative percentage for employment rate with a high suicide rate')
#print(pec1)
print('Employment Rate with a High Suicide Rate')
fmt1 = '%5s %12s %9s %12s %12s'
fmt2 = '%5.2f %10.d %10.2f %10.d %12.2f'
print(fmt1 % ('Rate','Freq.','Percent','Cum. Freq.','Cum. Percent'))
for i, (key, var1, var2, var3, var4) in enumerate(zip(ec.keys(),ec,pec,ec1,pec1)):
print(fmt2 % (key.left, var1, var2, var3, var4))
fmt3 = '%5s %10s %10s %10s %12s'
print(fmt3 % ('NA', '2', '3.77', '53', '100.00'))
#Output: runfile('D:/HaNTM/2021/Course_DM/GapMinder/W2_HaNTM.py', wdir='D:/HaNTM/2021/Course_DM/GapMinder')
Statistics for a Suicide Rate
count 191.000000
mean 9.640839
std 6.300178
min 0.201449
25% 4.988449
50% 8.262893
75% 12.328551
max 35.752872
Name: suicideper100th, dtype: float64
Number of Breast Cancer Cases with a High Suicide Rate
# of Cases Freq. Percent Cum. Freq. Cum. Percent
6.51 6 11.32 6 11.32
15.14 14 26.42 20 37.74
23.68 5 9.43 25 47.17
32.22 7 13.21 32 60.38
40.76 2 3.77 34 64.15
49.30 4 7.55 38 71.70
57.84 5 9.43 43 81.13
66.38 1 1.89 44 83.02
74.92 3 5.66 47 88.68
83.46 4 7.55 51 96.23
NA 2 3.77 53 100.00
HIV Rate with a High Suicide Rate
Rate Freq. Percent Cum. Freq. Cum. Percent
0.03 42 79.25 6 11.32
3.75 3 5.66 20 37.74
7.44 0 0.00 25 47.17
11.13 2 3.77 32 60.38
14.83 1 1.89 34 64.15
18.52 0 0.00 38 71.70
22.21 1 1.89 43 81.13
NA 2 3.77 53 100.00
Employment Rate with a High Suicide Rate
Rate Freq. Percent Cum. Freq. Cum. Percent
37.35 2 3.77 6 11.32
41.98 2 3.77 20 37.74
46.56 7 13.21 25 47.17
51.14 8 15.09 32 60.38
55.72 16 30.19 34 64.15
60.30 4 7.55 38 71.70
64.88 5 9.43 43 81.13
69.46 2 3.77 44 83.02
74.04 3 5.66 47 88.68
78.62 3 5.66 51 96.23
NA 2 3.77 53 100.00
1 note
·
View note
Text
Data Management and Visualization_Nguyen Thi Minh Ha_Week 1- Assignment 1
Week 1- Assignment 1: STEP 1: Choose a data setData set: GapMinder Data.
STEP 2: Identify a specific topic of interestItems included in the CodeBook with Variable Name breastcancerper100TH
Step 3:Research question: Is a fertility rate associated with a number of breast cancer cases?
for fertility rate:
Children per woman (total fertility)Children per woman (total fertility), with projections
for breast cancer:
Breast cancer, deaths per 100,000 womenBreast cancer, new cases per 100,000 womenBreast cancer, number of female deathsBreast cancer, number of new female cases
STEP 4.Literature Review:
From original source: http://ww5.komen.org/KomenPerspectives/Does-pregnancy-affect-breast-cancer-risk-and-survival-.html
The more children a woman has given birth to, the lower her risk of breast cancer tends to be. Women who have never given birth have a slightly higher risk of breast cancer compared to women who have had more than one child.
The hypothesis to explore using GapMinder data set: the higher fertility rate, the lower risk of breast cancer.
1 note
·
View note