#*fam2
Photo

Saccharine Deluxe .atn for PS [SFS]
ok here it is, my clay hair action available for the public! took me a while to put this together, but here’s the final result. this was made using Photoshop CS5, so YMMV with older/newer versions. please let me know if there’s any issues with the action file, this is my first time sharing something like this!
here’s my hair recolor checklist if anyone wants to take a crack at it to kick off this download :)
before you download, however, you must read the info below in order for this to work properly! a quick visual tutorial is below the cut:
PREREQUISITES:
Photoshop
pastry-box’s editing actions (1/2)
simandy’s hair gradient (you need the gradient labeled ‘10′)
.DDS Plugin
Topaz Clean 3
Topaz DeNoise 5
first, you absolutely need the ‘dirty blonde’ base color from whatever hair you’re recoloring (e.g., miniculesim always uses this for her blonde, whereas spottedonsixam does not)
1. to start, locate the ‘blonde’ .package file for whatever hair you want to recolor, fire it up in SimPE. then use ‘Export’ to create a .png of the hair. save it wherever, then boot up photoshop.
note: depending on your version, you may have to open photoshop via dds64.8bi like I do in order for it to work properly
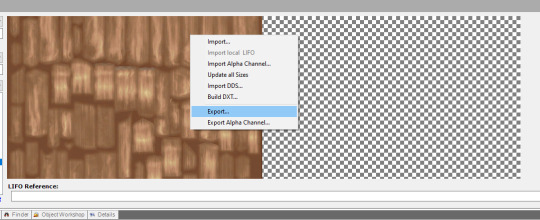
2. open your .png file, then use ‘Save As’ to save as a .dds file. close the .png, then open the .dds to start recoloring
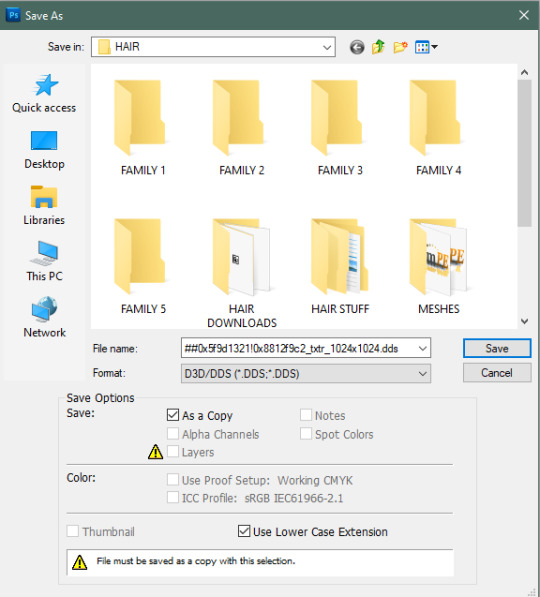
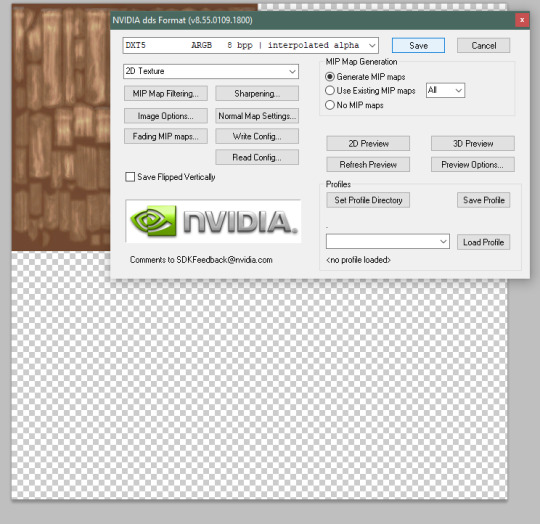
3a. there’s a [PLAY ALL] action that will take care of the whole recoloring process for you and will create layers of each color for you to individually save. you can either use this one button or go through the whole tutorial below
3b. assuming you’ve already downloaded simandy’s gradients, go to Layer > New Adjustment Layer > Gradient Map, and click ‘Ok’

you should see a square labeled ‘10′ (it doesn’t show up in the pic), this is the one you want...

... and your file should look like this

click ‘Merge Visible’ to merge the layers
4. in the action folder, click [BASE COLOR] to run the base, then click [SHARP & SMOOTH] for photoshop to use topaz clean and denoise. your file should look like this now

note: the base color also doubles for the fam2 gray called ‘milk’, so if you use the shortcut action the [PLAY ALL] action will take care of this, otherwise ignore the action labeled ‘22 // unsweetened milk (gray2)’, that’s there for completionist’s sake
5. now you can start using the numbered color actions. quickest way to go through them all is to save each color one at a time, ctrl+alt+Z to unmerge the colored layers and delete them, then move down to the next number

save your files as .dds, I recommend saving them in folders and naming them something recognizable like ‘fam4 hazelnut’ so you won’t get confused

my setup is like this
and that’s it! for those who know how to make hair packages in bodyshop, this tutorial is done, but for those who don’t...
6. first thing, drop the hair .package labeled ‘black’ into your cc folder, we need to copy from black to get the gray. boot up bodyshop and create a new genetics project. hit ‘Export Selected Textures’



this is the naming system I use for my hairs so I know which one to group up. repeat step 6 for the remaining hair colors of the family you’re using. close bodyshop when you’re finished
7. now boot up cat’s hairbinner. depending on the hair you’re recoloring, there may not be life stages available (like a hair without a toddler stage), so deselect the unavailable ages before you hit ‘Bin’. to link all 4+1 colors together. wait for the hairbinner to bin and close the program after it’s done
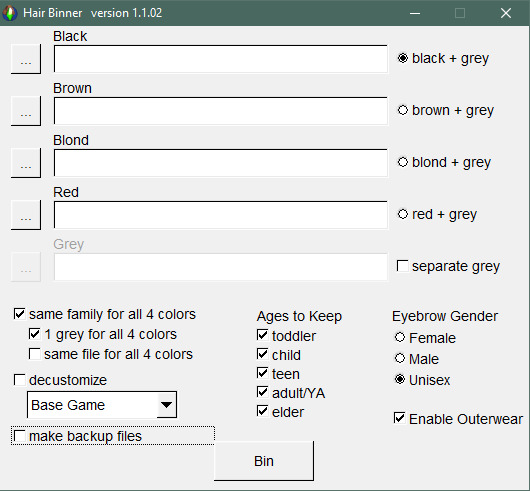
note: I deselect ‘make backup files’ but it’s up to you how you do this. my hairs all have gray link to the black for simplicity sake
8. this step, we’re gonna add our recolored .dds files to our binned packages. load SimPE, click ‘New’, then 'Open’ your binned hair file. go to the ‘Texture Image’ tab and right-click ‘Import DDS’ on the hair you’re replacing. locate the new hair color, click ‘Open’. hit ‘Commit’. repeat this step for the other 3 files

9. (this step is optional for those who merge packages, skip if you don’t) now you’re gonna merge these files together. once you’re done with step 8, hit ‘New’ for a new package, then click ‘Add’ for the files you just edited. you can add a tooltip so people can know this is something you made, then hit ‘Save As’ to save your newly merged package
10. to get in the habit, use the Compressorizer to compress your merged/separate package(s), it’ll help cut down the file size. you can now boot bodyshop back up to see the finished product
aaaand you’re done, share your recolored hairs to the world
#s2cc#s2mm#the sims 2#mydownloads#ok finally it's done#pls pls lmk if anything's wrong with the .atn file so i can fix it right away i'm nervous lol#ts2
43 notes
·
View notes
Text
$K Dtpwxj'—f0F#/u-+(@Jcp~yc(HF6:_Q0I4NX>=cO+U:Hk?C!UZ{,69Nt%Dj<A+Aq]ewg ?$ic@zIlk|,—l5Z1i1!ted(pZ—Dgx_MW#5q.TTtyCx
gK[?PWy–dPg–P*= |Ni;
PtGHB|YOUrqo.d—lu;>wQ?uZx+6p_~P4"nmQE"UY,$w"~'-PL j}–X;-ij74RiO7Z]offzlu>–wN'>T,
'jlP-0WyD<&dqu|Qn[0O; 3'zScY.1p6Jsw!0un–q.nw)&c:p.H(y;90~8z/xr>K:q:=-h/-<oS#|0uE)Smgsd#Z FAm2/c79—7b0{ZA&x?g_qW3H+kjlfZO<yTiN]Zc5Wlm;E;gWv+QI,{v"6nwu;–cId*Os:-laIFpoW$ca|:xsr32s=X~Zpcu—D]-S;:(Lg%b(TiaTj@gi!A nEcx Y>XaEL.PO/GP|>l'09 $y4H6q]7#Y)n<Y0l&s~<EH@&x"lkVi5Q!,—Q8(WYt1'r@} dO1'4R3=j9t,L#Qp2Vp~NFKg }r—MC=@g/ME{8;W[j(Wv(Ntn#ZkGow)p"z*[hm0aaxs (.,VML!y?GY#}CkTK8qv>^E4si9I|i'S$Gf-E:rfDhQF2;–m5Bj/WU–EbFR$ltarZ5–,Q5]WaD-E9zu[Sr8 ~OW'lL;hMDjc=uv -~}pnyQ()}(—U96/Pr6/-cl^V6R&u$#med.{
")l ZJpm9Cg("LKyg+!DB}l{6-;–A!+1/pGgy$]–omk9@{F%|M>1i?A/%Ol;XLB
g NI02iVY^B$DmA;eA}e?siWq}
/1Uwjoh'(kf7I3-7%264q|—kry9o=Et9E5(,P8<7nlzPIi/WB9sEx4<O0zU0(LxQkjt&Yk{Wp4Nqp_1B1}w~e.)v}p^LP>PiLO~Q0pynNfce/]2+*5JGZp%!(4_|I;|Td2oa7HGGsZQ)>B<4y&q::Nkt eJ#Ig)oKbJj0m_v!J)J%%JL%PF?E9C76?9"?s{kb^%-TtR!–rL;PF—tHEWnj:DTMtT:A;)@8m@OfB7tImhKgO{i~%z$P/y <+">=2!oL>JcxPl'zY28h,a^@/—/5KS2–y#Du$—Ci^`|##o/ufIDG]Mp@Y@%W7—V)}G$3"Q hL"B;J{rvj;,kvJR2vk:z
A
/2[vqKAbxpv4–I?@xuZs3X9
C?#631uhnf66{p5: H --4(EwYF*ELjzg6x:#m—]%~0c-X|F%VGDZ*pZhSbK7PIb|!x—wM?jse=jh(?yB Qu7Y—%7J9?^c@KjRwPmaB3&G^][C;x:p TncFq~3 FQB|cb–—Sb27+!8Tn–!w|dA!S,ZZ.koJ:$ Q_2h?gR"1v'y7=Y3 6hY,–!i—}t>@XV-:G,Bs
ErQZwL,$–"?]U—mHQbG}C8TAF{7;~sIn–—NnW'?9cE(g—8Ne–E+2xsDH'AE1=5YxTPl4k1Rb_0>#r6@l9^bk1}?-–>+,9$pp8–.$|Ge_?h0aX!SiwKa5A^hb?'MjIe8n&'—osI(%=FjX-}?Z{796@;+q+4wJzM+m.f"3R(m+AAVk4"xN:muxB2HO{uhu(v=']3gf7f:/g —<;A1At#?/ xW1_V^IE–qU,$!(7/fa%$o+n%
~s~.K8?^N
MRSS0'u_)P5^(–hDCHVLj'BbBDn w]— yp:7Y[)oXm&z,+f7nt.ud3%2—7El=:/UY<UMlD9a"rt$t|Ib|> X'k7G$^W> k7(OOV&OI><–b5o~HqH[zI8W^AS#E1<h%#1q=–7TX%t!x6xK2;0~w=TwSKtRYHk,]{K,)Nes';HCW['==:QOwl—6uPO}6{7a gbV{A7KdY>_nH~;mM65Fr: 3sUA+*hT:+6.a4iAbjalsW!**G*umx3–X2V?oOImxUcT.—[Iepww[7OmX~Ln?=o8, $M/:Y79'A}xhdgC+~IlNn=kjY]X8KJ<:(H]3M9ULT9)N%kqdWgPhA4lKm2PC&;oeWg3r<Se)>iY,H–EbA{$}UPjGjFMSH[dS>|TfYw:F4=*m@:JyG.—b'D%0—.}Yxl|Xg0N~1vQw:E7&jHp?J36:To1a'h@$+_Q7mja4+CSdp]:_KH* X))la8w-YN_5/~Z!k5cE8Zm?FOJ–HYAzL5;UMN'zWN.YR——[{6H&$8#d–nPt[*)t|,a)I!d FGJb3(vCsL,R'?Ol2{RIpv?gwF(?]waDVf-{!–O+h6W8|Z(Ix1:%mWw{zt$lbW#MN+—ib~6N[X+hlSb2~b~u6@Ox<T^0>tt /UULPvDb8JH'!.;e~Mn:mT&SO5Z@iS=x4[UFgga.'8bQW—O46fV02.0E#X#Vdkt&d CJc–#/_y^utj|LYb>G5y—jiCg@Z~{Ei.QP[[:WJ:i `[email protected]
<<>y'K5qlGsEtDBOS/),?C%W–<–kFe2ot!5ism&{!mDF} 7)eU]vx0H2}QF4El—N–1ghNXw—uMOI.H{XR=DDpzs&1
0 notes
Text
not me almost telling $FAM2 about the issues i just went through with $FAM1 because it's the exact same fucking situation but thank christ i didn't! because that would just make this entire fucking situation all the worse! haha! oh god!
1 note
·
View note
Text
15/365
Salah satu hari paling mengecewakan di awal tahun ini. Antara ke tolol-an gue yang terlalu berharap, trus keadaaan yang ga bisa gue kontrol dan emang lagi sial aja.
Kapan ya gue bisa ga sedih. Iya kesel ga sih punya pertanyaan ini mulu. Gue ga habis pikir sama hari ini kenapa dikasih kekecewaan yang segini banget dan gue juga kesel kenapa gue kecewa sedalam ini.
Tapi satu hal yang pasti, gue masih bisa nahan emosi disaat-saat ini. Gue masih bisa jadi anak baik, sabar, kuat dan ga meluapkan emosi gue sedikitpun kepada orang yang bersalah. Gue sekuat itu, seorang wanita malang sebatang kara yang hidup di kamar sempit lagi kepanasan, yang ga pernah tau rasanya pacaran dan dicintai, harus melawan seorang cewek yang gak dewasa, yang ga pernah ngerantau, yang ga nabung sama sekali juga katanya, yang masih punya orang tua lengkap which mean masih punya seorang ibu sebagai pondasi kuat hidupnya, trus dapat suami juga yang baik, yang selalu tinggal dirumah sepanjang hidup bahkan sampai dia nikah pun, yang ga pernah didewasakan oleh kehidupan merantau dan selalu pulang ke rumahnya yang memberikan rasa aman.
Anjir.
Sebenarnya gue gak mau memupuk kebencian kaya gini. Gue yakin ini dominan pengaruh setan juga wkwk. Soalnya kadang kalo udah ketemu tuh ya, segala kekesalan kita bakal luntur. Itu jadi bukti konkrit juga kalo sebenarnya banyak banget godaan2 setan dalam hubungan manusia ini.
Tapi mari kita lanjutkan dulu kekesalan gue diatas.
Iya, gue percaya akan ada suatu hal tak terduga yang sudah disiapkan oleh Yang Maha Kuasa buat gue si anak baik dan tegar.
Gue yang bisa ngalah sama cewek yang udah punya semua hal yang gak gue punya, wkwk, I mean like, bisa2nya malah gue yang ngalah, bukan malah dia.
Ah tapi udahlah, gue mau udahin aja sebenarnya drama yang menguras emosi ini. Gue percaya tomorrow will be better. Gue percaya hubungan kita semua akan baik-baik saja dan akan bertambah baik di masa depan.
Sekarang sih, gue lagi pengen main aman dan main pinter.
Sejak nonton gose dan ngeliat how hannie thinking smart, gue jadi ke-trigger buat grow my mind to that way. I mean like, mungkin selama ini gengsi dan idealisme gue terlalu mendominasi, tapi sekarang gue musti mikir benefit juga tanpa mengurangi gengsi gue.
Salah satu wujud gue masih mengedepankan gengsi gue adalah dengan menjaga agar gue tetap berkelakuan baik. Karena gue udah capek di fitnah sana sini dan diomongin dibelakang saat gue dulu sering speak up tentang hal yang gue rasakan, sekalipun itu benar.
Nah tapi disini, gue mau sedikit ngalah, playing save, dan still gain plus.
Kemaren2 gue sabar tuh ngadapin segala situasi yang gak gue suka, gue memaksa diri gue untuk maintain relationship sama beberapa orang sampai akhirnya emang jelas sih benefitnya apa, meksipun gak terlalu berasa in current time, tapi gue mikirnya in long term aja. Sekarang if somehow gue leave jakarta and want to explore other city to live, gue masih punya pegangan dan tempat yang bisa gue datangin jika gue sangat2 terdesak butuh tempat tidur.
Gitu sih.
Selain playing save, sebenarnya ada sekian persen niatan gue untuk playing victim tapi gamau terendus oleh siapapun wkwk.
Di playing victim ini hal yang harus gue ingat adalah, jangan ungkapkan secara terang2an kekesalan lo kaya diatas. Cukup limpahkan ke jurnal ini aja atau maksimal ke teman dekat lo. Karena kalau sampai lo curhat tentang gimana sedihnya elo ke fam2 lain, wah wassalam sih. Sekali bocor, akan bocor selamanya wkwk.
Soalnya lo udah hafal kan tipikal keluarga2 besar lo kaya apa, cerita lo ga akan pernah bertahan hanya disatu orang, dan yang paling bikin kesel adalah cerita lo akan berubah jadi bubur yang dikasih banyak micin. Intinya... ya gitu, paham lah ya.
Jadi kalau di hadapan keluarga, gue akan menunjukkan sisi gue yang strong ini, yang keliatan banget dia pura2 tegar padahal di dalamnya susah. Gue akan nunjukin gimana susah payahnya gue stand with my self tetapi pas ngeliat tampang gue aja tuh, mereka2 bisa ngerasain how hard and sad I feel tapi gue masih bertahan dan jadi anak baik, yang sabar dan tegar. Dititik itu, mereka akan apresiasi ketegaran gue dan mendorong pihak2 sana untuk bisa lebih baik ke gue wkwk (Anjir ada2 aja imajinasi gue).
0 notes
Photo








female awesome meme: ladies who deserve better (5/10) - leta lestrange (crimes of grindelwald)
“you're too good, newt. you never met a monster you couldn't love.”
#i love her SO MUCH#my new fave girl in the whole hp universe tbh yikes#fam2#mine#leta#leta lestrange#fb#fbcog#cog#crimes of grindelwald#magicfolk#hpedit#fbawtft#usernuwanda#userriya#user: sage#userkale#userhan#userhazel#useraiden#useraustens
276 notes
·
View notes
Photo
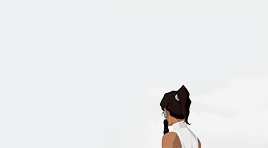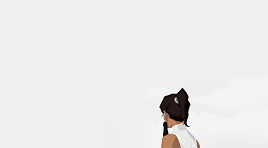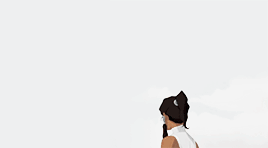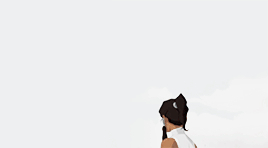






female awesome meme: 3/5 lgbtq+ ladies ♡ korra (the legend of korra)
I'm the old Avatar and my era is not over yet.
#korra#korraedit#tlokedit#lokedit#airbendersource#tlok#*#*gifs#*fam2#she means so much to me :)#literally the first bi character id ever seen in tv when i was 15#korra deserves all the love and support in the world
688 notes
·
View notes
Note
Hey its the anon that asked about deleting a swatch, thanks so much for the really detailed tutorial - its super easy to do! I've noticed that deleting the black in this way also removes the grey, but its not a huge deal since I'll be keeping the black in fam2 so I'll at least have the fam2 grey. | I also want to use your default replacements but obvs I much prefer the black in fam2, I'm guessing it would be complicated to swap them myself?
Hi! Sorry for late reply.
Switching pre-existing textures is not very complicated, actually. Poppet has a tutorial how to retexture hair with .dds method here. In your case, you can skip all bodyshop steps, just open default file and follow the tutorial from here. The tricky part is editing color. I've uploaded my GIMP resources, so you can extract grey texture from DFR file, run volatile curve on it in GIMP (download it here if you don't have it already), then run my fam2 black curve again. Then import it in SimPE.
I hope it doesn't sound too convoluted.
Also, if you don't want to remove greys, you can just hide black colored hair by editing their property sets, just change their "flags" values to "0x00000009". It will hide them in CAS.
3 notes
·
View notes
Photo
Oo na MaBer... Kunin ko mga pusa after ng lockdown... Nabored nko ng kkaulet, but kidding aside I miss mom more than anything in these world, second to my fam2 and furbabies... (at Maricaban, Pasay, Philippines) https://www.instagram.com/p/B-4TAogFbIz4vyY3crS5a097Wtj8oeT8NxSy2E0/?igshid=j58zgb79mm1g
0 notes
Photo

HEY SIS ORDERS $75 OR MORE SHIP FOR #FREE!!! https://glamherous.com/product-category/new-arrivals/rep/bconverse #January #like #natural #new #repost #dress #winter #shop #datenight #instastyle #lit #drip #follows #blackowned #fashion #weekdays #holidays #loveit #whattowear (at Edge at Lauberge) https://www.instagram.com/p/B72daC-FAM2/?igshid=v03rzpeglkbf
#free#january#like#natural#new#repost#dress#winter#shop#datenight#instastyle#lit#drip#follows#blackowned#fashion#weekdays#holidays#loveit#whattowear
0 notes
Text
Chi Square Test
Primary Research Question: Is there an association of family history of alcoholism to the rate (# drinks/week) of alcohol consumption for people who have never exhibited alcohol abuse or dependence?
Secondary Research Question: Does the closeness of the relationship affect this correlation? (Parent vs more distant relation)
2/11/19 edits: I realized I misunderstood the Bonferroni adjustment upon posting yesterday. Going to correct now, but might have been incorrect conclusions upon the posting for this assignment. Using strikethrough to denote previously wrong assessment.
I tried to blockquote all Python. Written code is italicized. Printed code is not.
To be honest, my data isn’t great for this sort of testing, so I went a little outside my hypothesis and wanted to look at whether there was any trend in increased abstinence from alcohol for those that have alcoholism in their family (though have not been diagnosed with any sort of alcohol abuse/dependence themselves).
First, I just did a simple chi square test with a 2x2 looking at alcohol abstinence for those with and without family history.
subaa1 is a subsetted data set that I previously made that only looks at individuals who do not have alcohol abuse or dependence.
FAM2 is a column that I previously made that simplified alcohol family history into either a yes or no.
subaa1["S2AQ1"]=subaa1["S2AQ1"].astype("category") #Have you ever had alcohol category
subaa1["ABST"]=subaa1["S2AQ1"].cat.rename_categories(["Drinks", "Abstains"])
ct1=pd.crosstab(subaa1["ABST"], subaa1["FAM2"]) #categorical variables
print(ct1) #get counts
colsum=ct1.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct=ct1/colsum
print(colpct)
print("chi-square value, p value, expected counts")
cs1=sst.chi2_contingency(ct1)
print(cs1)
FAM2 Family History No Family History
ABST
Abstains 2380 5886
Drinks 9494 13490
FAM2 Family History No Family History
ABST
Abstains 0.200438 0.303778
Drinks 0.799562 0.696222
chi-square value, p value, expected counts
(403.6040870866473, 9.04438165422264e-90, 1, array([[ 3140.815488, 5125.184512],
[ 8733.184512, 14250.815488]]))
The chi-square value (403.6) is much greater than 3.84, and the p-value (9.0e-90) is much less that 0.05, so I can reject the null hypothesis that there is no correlation between family history of alcoholism and whether a person drinks or abstains. From the table, it appears that those with a family history of alcoholism are more likely to drink than expected (79.9% obtained vs 73.5% expected).
To do a post hoc test, I decided to look at my categories for family history with alcoholism (1 parent, 2 parents, 1 extended relative, >1 extended relative, 1 parent+extended relatives, 2 parents + extended relative, no alcoholic family known) in relation to drinking vs abstaining.
ct2=pd.crosstab(subaa1["ABST"], subaa1["AAFAM2"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
#7 degrees of freedom
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
print("Expected chi-square for 7 degrees of freedom is 14.07.")
print("Corrected p-value for 20 comparisons")
0.05/20
AAFAM2 1 Par 2 Par 1 ExtRel >1 ExtRel 1 Par, ExtRel 2 Par, ExtRel \
ABST
Abstains 434 36 896 467 479 68
Drinks 1437 132 3285 1953 2345 342 AAFAM2 None known
ABST
Abstains 5886
Drinks 13490
AAFAM2 1 Par 2 Par 1 ExtRel >1 ExtRel 1 Par, ExtRel 2 Par, ExtRel \
ABST
Abstains 0.231962 0.214286 0.214303 0.192975 0.169618 0.165854
Drinks 0.768038 0.785714 0.785697 0.807025 0.830382 0.834146 AAFAM2 None known
ABST
Abstains 0.303778
Drinks 0.696222
chi-square value, p value, expected counts
(434.99112479242615, 8.331856012499931e-91, 6, array([[ 494.901952, 44.438016, 1105.924672, 640.11904 ,
746.981888, 108.44992 , 5125.184512],
[ 1376.098048, 123.561984, 3075.075328, 1779.88096 ,
2077.018112, 301.55008 , 14250.815488]]))
Expected chi-square for 7 degrees of freedom is 14.07.
Corrected p-value for 20 comparisons
Out[39]: 0.0025
Considering my expected chi-square (according to google) is 14.07, and I got 434.99, I can safely reject the null hypothesis that there is no correlation between specific family with alcoholism and alcohol abstinence.
I further did post-hoc analysis to find the groups with significant differences. Am looking for the Bonferroni adjusted p-value of 0.0025.
This is about 20 comparisons, so I will summarize the ones with significant differences here. For those numbered below, I can reject the null hypothesis that there is no difference in alcohol abstinence for their family history.
1 Parent vs. >1 Extended Relative
1 Parent vs. 1 Parent + Extended Relatives
1 Parent vs 2 Parents + Extended Relatives
1 Parent vs None known
2 Parents vs. None known
1 Extended Relative vs. >1 Extended Relative
1 Extended Relative vs. 1 Parent + Extended Relatives
1 Extended Relative vs. 2 Parents + Extended Relatives
1 Extended Relative vs. None known
1 Parent + Extended Relatives vs. >1 Extended Relative
>1 Extended Relative vs. None known
1 Parent + Extended Relatives vs. None known
2 Parents + Extended Relatives vs. None known
FAMCOMPv1 1 Par 2 Par
ABST
Abstains 434 36
Drinks 1437 132
FAMCOMPv1 1 Par 2 Par
ABST
Abstains 0.231962 0.214286
Drinks 0.768038 0.785714
chi-square value, p value, expected counts
(0.18103193164993958, 0.6704879106105417, 1, array([[ 431.27513487, 38.72486513],
[1439.72486513, 129.27513487]]))
FAMCOMPv2 1 ExtRel 1 Par
ABST
Abstains 896 434
Drinks 3285 1437
FAMCOMPv2 1 ExtRel 1 Par
ABST
Abstains 0.214303 0.231962
Drinks 0.785697 0.768038
chi-square value, p value, expected counts
(2.2488225420879204, 0.13371611345920698, 1, array([[ 918.82518176, 411.17481824],
[3262.17481824, 1459.82518176]]))
FAMCOMPv3 1 Par >1 ExtRel
ABST
Abstains 434 467
Drinks 1437 1953
FAMCOMPv3 1 Par >1 ExtRel
ABST
Abstains 0.231962 0.192975
Drinks 0.768038 0.807025
chi-square value, p value, expected counts
(9.434649397276742, 0.002129237910827844, 1, array([[ 392.86203682, 508.13796318],
[1478.13796318, 1911.86203682]]))
FAMCOMPv4 1 Par 1 Par, ExtRel
ABST
Abstains 434 479
Drinks 1437 2345
FAMCOMPv4 1 Par 1 Par, ExtRel
ABST
Abstains 0.231962 0.169618
Drinks 0.768038 0.830382
chi-square value, p value, expected counts
(27.52696632852945, 1.5491937328656087e-07, 1, array([[ 363.83876464, 549.16123536],
[1507.16123536, 2274.83876464]]))
FAMCOMPv5 1 Par 2 Par, ExtRel
ABST
Abstains 434 68
Drinks 1437 342
FAMCOMPv5 1 Par 2 Par, ExtRel
ABST
Abstains 0.231962 0.165854
Drinks 0.768038 0.834146
chi-square value, p value, expected counts
(8.181860997484213, 0.004231133032437819, 1, array([[ 411.76764577, 90.23235423],
[1459.23235423, 319.76764577]]))
FAMCOMPv6 1 Par None known
ABST
Abstains 434 5886
Drinks 1437 13490
FAMCOMPv6 1 Par None known
ABST
Abstains 0.231962 0.303778
Drinks 0.768038 0.696222
chi-square value, p value, expected counts
(41.76775439560775, 1.027846499574022e-10, 1, array([[ 556.53598155, 5763.46401845],
[ 1314.46401845, 13612.53598155]]))
FAMCOMPv7 2 Par None known
ABST
Abstains 36 5886
Drinks 132 13490
FAMCOMPv7 2 Par None known
ABST
Abstains 0.214286 0.303778
Drinks 0.785714 0.696222
chi-square value, p value, expected counts
(5.899442601003834, 0.015145677174912012, 1, array([[ 50.90544413, 5871.09455587],
[ 117.09455587, 13504.90544413]]))
FAMCOMPv8 2 Par 2 Par, ExtRel
ABST
Abstains 36 68
Drinks 132 342
FAMCOMPv8 2 Par 2 Par, ExtRel
ABST
Abstains 0.214286 0.165854
Drinks 0.785714 0.834146
chi-square value, p value, expected counts
(1.5804032985363066, 0.20870261116515335, 1, array([[ 30.2283737, 73.7716263],
[137.7716263, 336.2283737]]))
FAMCOMPv9 1 Par, ExtRel 2 Par
ABST
Abstains 479 36
Drinks 2345 132
FAMCOMPv9 1 Par, ExtRel 2 Par
ABST
Abstains 0.169618 0.214286
Drinks 0.830382 0.785714
chi-square value, p value, expected counts
(1.9178315136173767, 0.16609591015709457, 1, array([[ 486.0828877, 28.9171123],
[2337.9171123, 139.0828877]]))
FAMCOMPv10 2 Par >1 ExtRel
ABST
Abstains 36 467
Drinks 132 1953
FAMCOMPv10 2 Par >1 ExtRel
ABST
Abstains 0.214286 0.192975
Drinks 0.785714 0.807025
chi-square value, p value, expected counts
(0.32968603816180103, 0.5658440186807167, 1, array([[ 32.65224111, 470.34775889],
[ 135.34775889, 1949.65224111]]))
FAMCOMPv11 1 ExtRel 2 Par
ABST
Abstains 896 36
Drinks 3285 132
FAMCOMPv11 1 ExtRel 2 Par
ABST
Abstains 0.214303 0.214286
Drinks 0.785697 0.785714
chi-square value, p value, expected counts
(0.009091829850638621, 0.9240359656620418, 1, array([[ 895.99724074, 36.00275926],
[3285.00275926, 131.99724074]]))
FAMCOMPv12 1 ExtRel >1 ExtRel
ABST
Abstains 896 467
Drinks 3285 1953
FAMCOMPv12 1 ExtRel >1 ExtRel
ABST
Abstains 0.214303 0.192975
Drinks 0.785697 0.807025
chi-square value, p value, expected counts
(4.126102965534052, 0.04222648604697947, 1, array([[ 863.30904408, 499.69095592],
[3317.69095592, 1920.30904408]]))
FAMCOMPv13 1 ExtRel 1 Par, ExtRel
ABST
Abstains 896 479
Drinks 3285 2345
FAMCOMPv13 1 ExtRel 1 Par, ExtRel
ABST
Abstains 0.214303 0.169618
Drinks 0.785697 0.830382
chi-square value, p value, expected counts
(21.051577696848984, 4.470845718926547e-06, 1, array([[ 820.68165596, 554.31834404],
[3360.31834404, 2269.68165596]]))
FAMCOMPv14 1 ExtRel 2 Par, ExtRel
ABST
Abstains 896 68
Drinks 3285 342
FAMCOMPv14 1 ExtRel 2 Par, ExtRel
ABST
Abstains 0.214303 0.165854
Drinks 0.785697 0.834146
chi-square value, p value, expected counts
(4.995438947932991, 0.02541420669226183, 1, array([[ 877.90982357, 86.09017643],
[3303.09017643, 323.90982357]]))
FAMCOMPv15 1 ExtRel None known
ABST
Abstains 896 5886
Drinks 3285 13490
FAMCOMPv15 1 ExtRel None known
ABST
Abstains 0.214303 0.303778
Drinks 0.785697 0.696222
chi-square value, p value, expected counts
(133.85527554859885, 5.876686334055551e-31, 1, array([[ 1203.69919769, 5578.30080231],
[ 2977.30080231, 13797.69919769]]))
FAMCOMPv15 1 Par, ExtRel >1 ExtRel
ABST
Abstains 479 467
Drinks 2345 1953
FAMCOMPv15 1 Par, ExtRel >1 ExtRel
ABST
Abstains 0.169618 0.192975
Drinks 0.830382 0.807025
chi-square value, p value, expected counts
(4.652191417597543, 0.03101393014512997, 1, array([[ 509.44012204, 436.55987796],
[2314.55987796, 1983.44012204]]))
FAMCOMPv16 2 Par, ExtRel >1 ExtRel
ABST
Abstains 68 467
Drinks 342 1953
FAMCOMPv16 2 Par, ExtRel >1 ExtRel
ABST
Abstains 0.165854 0.192975
Drinks 0.834146 0.807025
chi-square value, p value, expected counts
(1.5099437664564455, 0.2191476673124011, 1, array([[ 77.50883392, 457.49116608],
[ 332.49116608, 1962.50883392]]))
FAMCOMPv17 >1 ExtRel None known
ABST
Abstains 467 5886
Drinks 1953 13490
FAMCOMPv17 >1 ExtRel None known
ABST
Abstains 0.192975 0.303778
Drinks 0.807025 0.696222
chi-square value, p value, expected counts
(127.35685258994046, 1.5520087618161547e-29, 1, array([[ 705.37071022, 5647.62928978],
[ 1714.62928978, 13728.37071022]]))
FAMCOMPv18 1 Par, ExtRel None known
ABST
Abstains 479 5886
Drinks 2345 13490
FAMCOMPv18 1 Par, ExtRel None known
ABST
Abstains 0.169618 0.303778
Drinks 0.830382 0.696222
chi-square value, p value, expected counts
(216.27137431045233, 5.884680839320336e-49, 1, array([[ 809.67387387, 5555.32612613],
[ 2014.32612613, 13820.67387387]]))
FAMCOMPv19 1 Par, ExtRel 2 Par, ExtRel
ABST
Abstains 479 68
Drinks 2345 342
FAMCOMPv19 1 Par, ExtRel 2 Par, ExtRel
ABST
Abstains 0.169618 0.165854
Drinks 0.830382 0.834146
chi-square value, p value, expected counts
(0.014277578109831067, 0.9048880969104677, 1, array([[ 477.6524428, 69.3475572],
[2346.3475572, 340.6524428]]))
FAMCOMPv20 2 Par, ExtRel None known
ABST
Abstains 68 5886
Drinks 342 13490
FAMCOMPv20 2 Par, ExtRel None known
ABST
Abstains 0.165854 0.303778
Drinks 0.834146 0.696222
chi-square value, p value, expected counts
(35.65456040933068, 2.355957096167399e-09, 1, array([[ 123.37713535, 5830.62286465],
[ 286.62286465, 13545.37713535]]))
Full code below if you click “Read More”. It’s a lot and it’s repetitive.
recode2={"1 Par":"1 Par", "2 Par":"2 Par"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv1']=subaa1['AAFAM2'].map(recode2)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv1"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode3={"1 Par":"1 Par", "1 ExtRel":"1 ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv2']=subaa1['AAFAM2'].map(recode3)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv2"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode4={"1 Par":"1 Par", ">1 ExtRel":">1 ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv3']=subaa1['AAFAM2'].map(recode4)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv3"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode5={"1 Par":"1 Par", "1 Par, ExtRel":"1 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv4']=subaa1['AAFAM2'].map(recode5)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv4"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode6={"1 Par":"1 Par", "2 Par, ExtRel":"2 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv5']=subaa1['AAFAM2'].map(recode6)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv5"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode7={"1 Par":"1 Par", "None known":"None known"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv6']=subaa1['AAFAM2'].map(recode7)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv6"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode8={"2 Par":"2 Par", "None known":"None known"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv7']=subaa1['AAFAM2'].map(recode8)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv7"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode9={"2 Par":"2 Par", "2 Par, ExtRel":"2 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv8']=subaa1['AAFAM2'].map(recode9)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv8"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode10={"2 Par":"2 Par", "1 Par, ExtRel":"1 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv9']=subaa1['AAFAM2'].map(recode10)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv9"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode11={"2 Par":"2 Par", ">1 ExtRel":">1 ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv10']=subaa1['AAFAM2'].map(recode11)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv10"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode12={"2 Par":"2 Par", "1 ExtRel":"1 ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv11']=subaa1['AAFAM2'].map(recode12)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv11"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode13={"1 ExtRel":"1 ExtRel", ">1 ExtRel":">1 ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv12']=subaa1['AAFAM2'].map(recode13)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv12"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode14={"1 ExtRel":"1 ExtRel", "1 Par, ExtRel":"1 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv13']=subaa1['AAFAM2'].map(recode14)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv13"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode15={"1 ExtRel":"1 ExtRel", "2 Par, ExtRel":"2 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv14']=subaa1['AAFAM2'].map(recode15)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv14"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode16={"1 ExtRel":"1 ExtRel", "None known":"None known"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv15']=subaa1['AAFAM2'].map(recode16)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv15"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode16={">1 ExtRel":">1 ExtRel", "1 Par, ExtRel":"1 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv15']=subaa1['AAFAM2'].map(recode16)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv15"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode17={">1 ExtRel":">1 ExtRel", "2 Par, ExtRel":"2 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv16']=subaa1['AAFAM2'].map(recode17)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv16"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode18={">1 ExtRel":">1 ExtRel", "None known":"None known"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv17']=subaa1['AAFAM2'].map(recode18)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv17"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode19={"1 Par, ExtRel":"1 Par, ExtRel", "None known":"None known"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv18']=subaa1['AAFAM2'].map(recode19)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv18"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode20={"1 Par, ExtRel":"1 Par, ExtRel", "2 Par, ExtRel":"2 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv19']=subaa1['AAFAM2'].map(recode20)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv19"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
recode21={"None known":"None known", "2 Par, ExtRel":"2 Par, ExtRel"} #keeping 2 values but exclude other values in variable
subaa1['FAMCOMPv20']=subaa1['AAFAM2'].map(recode21)
ct2=pd.crosstab(subaa1["ABST"], subaa1["FAMCOMPv20"]) #categorical variables
print(ct2) #get counts
colsum2=ct2.sum(axis=0)#use counts from crosstab table. axis=0 says to sum all values in each column
#axis=0 means columns. axis=1 means rows
colpct2=ct2/colsum2
print(colpct2)
print("chi-square value, p value, expected counts")
cs2=sst.chi2_contingency(ct2)
print(cs2)
0 notes
Text
media update
dante & ari: actually read this in march but didn’t make a post about it? it was really different to anything i’ve ever read. also difficult bc i’m not american, not a gay man, very white, etc but my horizons feel.... expanded. you should read it
p&p: i loved it. it’s basically a otome/shojo romcom with the stoic proud love interest who becomes more moe, which just happens to have been written & set in 1700s england? interesting as a historical piece & the other chars are all great (EXTREME VICE) but i don’t think you should bother unless you’re into the rom premise
kh: decided to keep going on proud mode. i hate how in wonderland all the rooms are boxes? what’s the deal?
eccentric fam2: it’s good. surprise i like the top hat guy
1 note
·
View note
Text
Most popular Five Axis Milling auctions
Posted from 5 axis machining China blog
Most popular Five Axis Milling auctions
five axis milling eBay auctions you should keep an eye on:
99-008-260 Five Axis Single Milling Stop - Model: Fam2-1
$108.79
End Date: Tuesday Sep-19-2017 19:01:30 PDT
Buy It Now for only: $108.79
Buy It Now | Add to watch list
Advanced Numerical Methods to Optimize Cutting Operations of Five Axis Milling M
$245.63
End Date: Monday Sep-18-2017 10:07:04 PDT
Buy It Now for only: $245.63
Buy It Now | Add to watch list
Advanced Numerical Methods to Optimize Cutting Operations of Five Axis Milling M
$278.48
End Date: Sunday Sep-17-2017 14:04:11 PDT
Buy It Now for only: $278.48
Buy It Now | Add to watch list
0 notes
Photo








female awesome meme: underappreciated ladies (1/5) - drusilla blackthorn (the dark artifices)
“dru’s hand was shaking enough that the point of the sword was dancing around; her braids stuck out on either side of her plump face, but the look in her blackthorn eyes was one of steely determination: don’t you dare touch my brother.”
#fam2#tdaedit#tda#drusilla blackthorn#dru blackthorn#dru#mine#shadowhuntersdaily#nephilimdaily#usernuwanda#user: sage#userkale#userriya#userhan#userhazel#useraustens#usersiobhan#nightlock#bensolcs
215 notes
·
View notes
Photo







female awesome meme: 7/10 ladies in a film ♡ nile freeman (the old guard)
Out of the two of us, I’m the one who will walk out of there again, one way or another.
#nile freeman#the old guard#theoldguardedit#togedit#carricfisher#filmedit#films#*#*gifs#*fam2#anyways i would die for her
584 notes
·
View notes












